

Molecular Modelling for Immunology
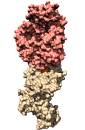
Web site for the 3D Structure & Modelling workshop of the Master in Medical Biology, Immunology And Cancer, from the University of Lausanne.
This web site has been designed to be used with the Chrome web browser. Please install Chrome to use this web page.
Several links are accessible on this page that allow visualizing 3D structures of proteins on computers equiped with the UCSF Chimera program. To use this web page, please, install the "current production release" version of UCSF Chimera that you can find here: Download UCSF Chimera.
Note that all machines of the B304 room of Epalinges are already setup correctly. No further installation is required and 3D structures will be automatically loaded in UCSF chimera after clicking on the links.
This web site has been designed to be used with the Chrome web browser. Please install Chrome to use this web page.
Several links are accessible on this page that allow visualizing 3D structures of proteins on computers equiped with the UCSF Chimera program. To use this web page, please, install the "current production release" version of UCSF Chimera that you can find here: Download UCSF Chimera.
Note that all machines of the B304 room of Epalinges are already setup correctly. No further installation is required and 3D structures will be automatically loaded in UCSF chimera after clicking on the links.
Lecture
The PDF file of the lecture's slides can be found here.
The following links give you access to the UCSF Chimera 3D sessions corresponding to the lecture's illustrations. This will allow you to manipulate in 3D the structures that will be displayed during the lecture.
Prologue: molecular representations
Origin of the 3D structures
Molecular recognition
 Slide 43.
Overview of the interactions between HLA-A*0201 and MART-1/Melan-A.
Slide 43.
Overview of the interactions between HLA-A*0201 and MART-1/Melan-A. Slide 56.
Non Polar interactions.
Slide 56.
Non Polar interactions.You can change the representation of residue #2:2 to "sphere" for a better visualization of the space filling.
 Slide 60.
&
Slide 60.
Hydrogen bonds.
Slide 60.
&
Slide 60.
Hydrogen bonds. Slide 63.
π interactions.
Slide 63.
π interactions. Slide 65
&
Slide 65.
Role of water molecules.
Slide 65
&
Slide 65.
Role of water molecules. Slide 70.
Influence of flexibility and entropy.
Slide 70.
Influence of flexibility and entropy.3D structure of the peptide-MHC Class I complex.
 Slide 77
&
Slide 77.
Global structure of MHC Class I.
Slide 77
&
Slide 77.
Global structure of MHC Class I. Slide 78.
Secondary structure elements of the binding pocket.
Slide 78.
Secondary structure elements of the binding pocket. Slide 79
&
Slide 79.
"Walls" of the binding pocket that limits the size of peptide binders.
Slide 79
&
Slide 79.
"Walls" of the binding pocket that limits the size of peptide binders. Slide 80.
Hydrogen bonds at the N and C-termini.
Slide 80.
Hydrogen bonds at the N and C-termini. Slide 82 & Slide 83.
Peptide anchor residues in HLA-A*0201.
Slide 82 & Slide 83.
Peptide anchor residues in HLA-A*0201. Slide 84.
Peptide secondary anchor residues in HLA-A*0201 & Peptides residues facing the solvent or the TCR.
Slide 84.
Peptide secondary anchor residues in HLA-A*0201 & Peptides residues facing the solvent or the TCR.3D structure of the peptide-MHC Class II complex.
 Slide 89 & Slide 90.
Structure of pMHC Class II.
Slide 89 & Slide 90.
Structure of pMHC Class II.Open the "Model Panel" to show/hide surfaces.
3D structure of the complex between TCR and pMHC Class I.
 Slide 94.
Structure of the T-cell receptor (TCR).
Slide 94.
Structure of the T-cell receptor (TCR). Slide 95.
Structure of the TCR/pMHC complex.
Slide 95.
Structure of the TCR/pMHC complex.Open the "Model Panel" to show/hide surfaces.
 Slide 96.
Structure of the TCR/pMHC complex, bound to CD8.
Slide 96.
Structure of the TCR/pMHC complex, bound to CD8.
Exercises
The PDF file of the exercises can be found here.